Spatial Temporal Graph Convolution with Graph Structure Self-learning for Early MCI Detection
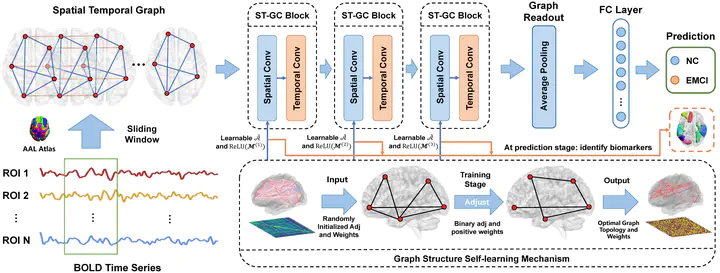 Framework Overview
Framework Overview
Abstract
Graph neural networks (GNNs) have been successfully applied to early mild cognitive impairment (EMCI) detection, with the usage of elaborately designed features constructed from blood oxygen level-dependent (BOLD) time series. However, few works explored the feasibility of using BOLD signals directly as features. Meanwhile, existing GNN-based methods primarily rely on hand-crafted explicit brain topology as the adjacency matrix, which is not optimal and ignores the implicit topological organization of the brain. In this paper, we propose a spatial temporal graph convolutional network with a novel graph structure self-learning mechanism for EMCI detection. The proposed spatial temporal graph convolution block directly exploits BOLD time series as input features, which provides an interesting view for rsfMRIbased preclinical AD diagnosis. Moreover, our model can adaptively learn the optimal topological structure and refine edge weights with the graph structure self-learning mechanism. Results on the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database show that our method outperforms state-of-the-art approaches. Biomarkers consistent with previous studies can be extracted from the model, proving the reliable interpretability of our method.
Supplementary notes can be added here, including code, math, and images.